Studying Weak and Dynamic Interactions of Posttranslationally Modified Proteins using Expressed Protein Ligation
03-Dec-2013
ACS Chem. Biol., 2013, DOI: 10.1021/cb400723j, 9 (2), pp 347–352 published on 03.12.2013
ACS Chem. Biol., online article
ACS Chem. Biol., online article
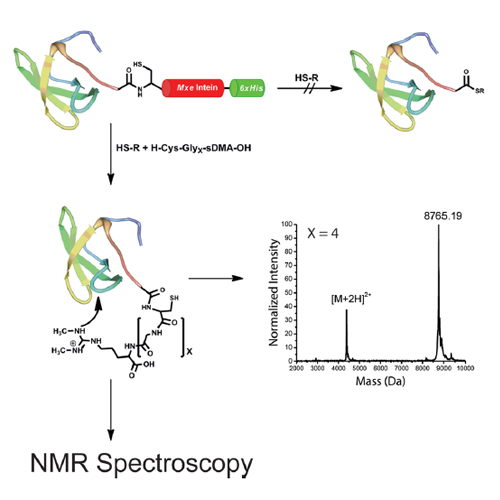
Many cellular processes are regulated by posttranslational modifications that are recognized by specific domains in protein binding partners. These interactions are often weak, thus allowing a highly dynamic and combinatorial regulatory network of protein–protein interactions. We report an efficient strategy that overcomes challenges in structural analysis of such a weak transient interaction between the Tudor domain of the Survival of Motor Neuron (SMN) protein and symmetrically dimethylated arginine (sDMA). The posttranslational modification is chemically introduced and covalently linked to the effector module by a one-pot expressed protein ligation (EPL) procedure also enabling segmental incorporation of NMR-active isotopes for structural analysis. Covalent coupling of the two interacting moieties shifts the equilibrium to the bound state, and stoichiometric interactions are formed even for low affinity interactions. Our approach should enable the structural analysis of weak interactions by NMR or X-ray crystallography to better understand the role of posttranslational modifications in dynamic biological processes.