Exploring Adenylylation and Phosphocholination as Post-Translational Modifications
03-Jan-2014
ChemBioChem, 2014, DOI: 10.1002/cbic.201300508, Volume 15, Issue 1, pages 19–26, published on 03.01.2014
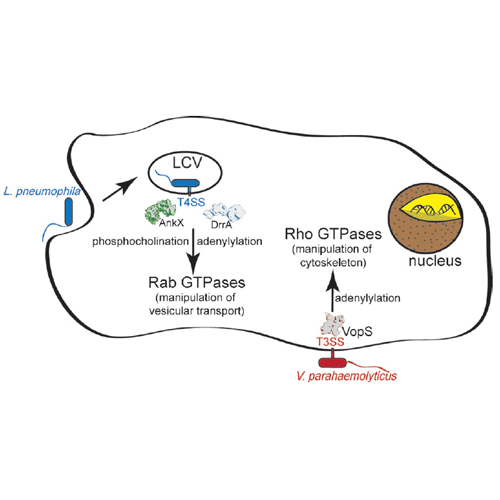
Pathogenic bacteria have developed a variety of mechanisms with which to manipulate the functions of proteins in important cellular pathways of host organisms. Because of their pivotal role in the regulation of cell physiology, members of the family of G-proteins and cytoskeletal proteins are frequent targets for pathogens. They are modified both by noncovalent interaction with bacterial virulence factors and by post-translational modification (PTMs; e.g., proteolysis, deamidation, transglutamination, ADP-ribosylation adenylylation, phosphocholination, glucosylation). This review focuses on covalent modification of target proteins by nucleotidylylation and phosphocholination by pathogens and their role in modulating host protein activity. The identification of proteins modified with these PTMs in vivo will give insights into the role of the modifications during the bacterial infection process. For this reason, special emphasis is put on chemical proteomics and on the development of tools for the in vivo detection and proteomic dissection of such PTMs (for recent progress in this field see refs and Introducing small chemical handles into the donor substrates potentially facilitates the identification of protein substrates modified by specific enzymes. This review discusses recent advances in the exploitation of such chemical modifications. Also, we describe the use of specific antibodies raised against modified proteins, peptides, and amino acids, as a complementary tool for the proteomic dissection of the PTMs.