In vivo aspects of protein folding and quality control
01-Jul-2016
Science, Vol. 353, Issue 6294, DOI: 10.1126/science.aac4354
Science, online article
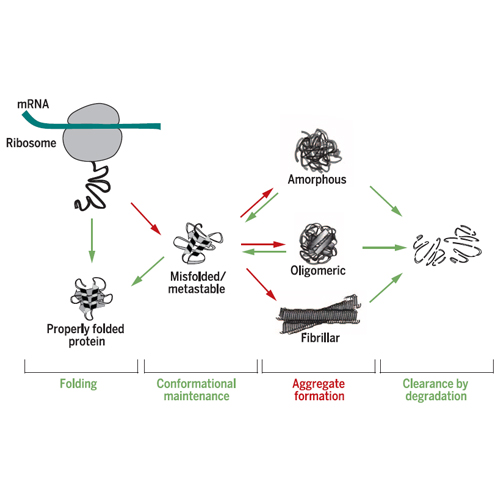
Proteins are synthesized on ribosomes as linear chains of amino acids and must fold into unique three-dimensional structures to fulfill their biological functions. Protein folding is intrinsically error-prone, and how it is accomplished efficiently represents a problem of great biological and medical importance. During folding, the nascent polypeptide must navigate a complex energy landscape. As a result, misfolded molecules may accumulate that expose hydrophobic amino acid residues and thus are in danger of forming potentially toxic aggregates. To ensure efficient folding and prevent aggregation, cells in all domains of life express various classes of proteins called molecular chaperones. These proteins receive the nascent polypeptide chain emerging from the ribosome and guide it along a productive folding pathway. Because proteins are structurally dynamic, constant surveillance of the proteome by an integrated network of chaperones and protein degradation machineries, the proteostasis network (PN), is required to maintain protein homeostasis in a range of external and endogenous stress conditions.