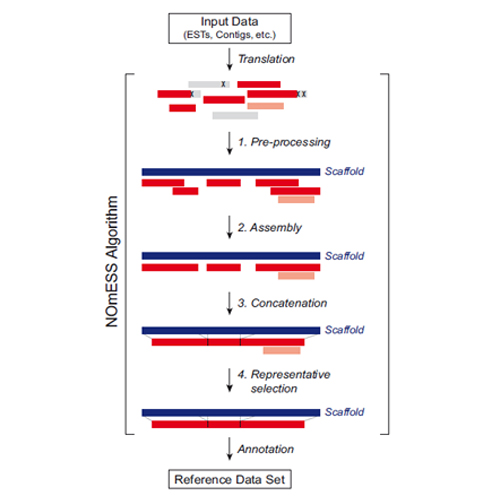
Homology-driven assembly of NOn-redundant protEin sequence sets (NOmESS) for mass spectrometry
06-Jan-2015
Bioinformatics, 32 (9): 1417-1419, DOI: https://doi.org/10.1093/bioinformatics/btv756
Bioinformatics, online article
Summary: To enable mass spectrometry (MS)-based proteomic studies with poorly characterized organisms, we developed a computational workflow for the homology-driven assembly of a non-redundant reference sequence dataset. In the automated pipeline, translated DNA sequences (e.g. ESTs, RNA deep-sequencing data) are aligned to those of a closely related and fully sequenced organism. Representative sequences are derived from each cluster and joined, resulting in a non-redundant reference set representing the maximal available amino acid sequence information for each protein. We here applied NOmESS to assemble a reference database for the widely used model organism Xenopus laevis and demonstrate its use in proteomic applications.