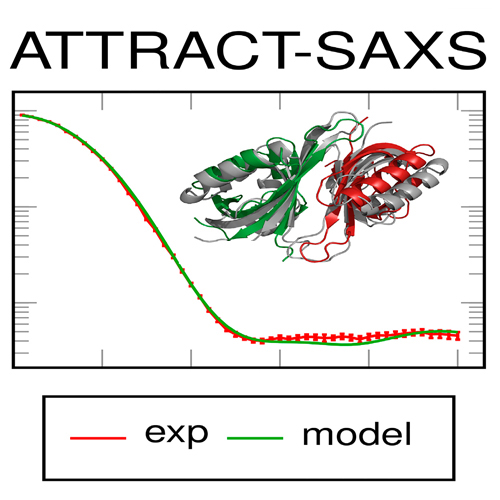
SAXS Data Alone can Generate High-Quality Models of Protein-Protein Complexes
02-Aug-2016
Structure, Volume 24, Issue 8, p1387–1397, http://dx.doi.org/10.1016/j.str.2016.06.007
Structure, online article
Modeling driven by small-angle X-ray scattering (SAXS) combines low-resolution data with computational modeling to predict the structure of biomolecular assemblies. A new protocol, ATTRACT-SAXS, has been developed and tested on a large protein-protein docking benchmark with simulated SAXS data. For 88% of cases, high-quality solutions were generated using SAXS data alone without a physiochemical force field (interface-RMSD ≦ 2 Å or ligand-RMSD ≦ 5 Å; and more than 30% native contacts). ATTRACT-SAXS gave significant improvements compared with previous approaches that filter by SAXS a posteriori. When combining SAXS and interface properties for scoring, the protocol placed high-quality models in 70% of cases among the top-ranked 100 clusters. ATTRACT-SAXS also gave good results when tested on experimental data if the native complex structure was compatible with the SAXS profile. Our results show that, in principle, SAXS on its own can contain enough information to generate high-quality models of protein-protein complexes.