MC EMiNEM Maps the Interaction Landscape of the Mediator
21-Jun-2012
PLoS Computational Biology, 2012, doi:10.1371/journal.pcbi.1002568, Volume 8 , Issue 6 published on 21.06.2012
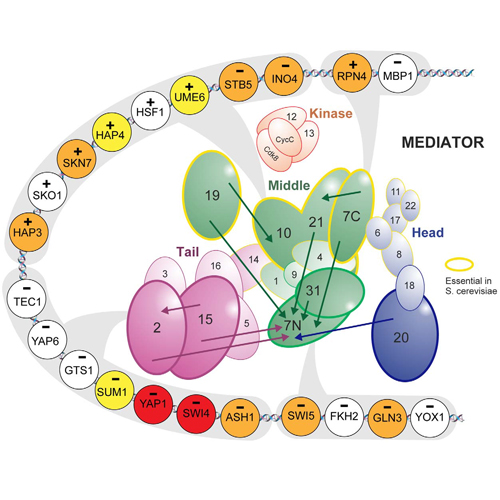
The Mediator is a highly conserved, large multiprotein complex that is involved essentially in the regulation of eukaryotic mRNA transcription. It acts as a general transcription factor by integrating regulatory signals from gene-specific activators or repressors to the RNA Polymerase II. The internal network of interactions between Mediator subunits that conveys these signals is largely unknown. Here, we introduce MC EMiNEM, a novel method for the retrieval of functional dependencies between proteins that have pleiotropic effects on mRNA transcription. MC EMiNEM is based on Nested Effects Models (NEMs), a class of probabilistic graphical models that extends the idea of hierarchical clustering. It combines mode-hopping Monte Carlo (MC) sampling with an Expectation-Maximization (EM) algorithm for NEMs to increase sensitivity compared to existing methods. A meta-analysis of four Mediator perturbation studies in Saccharomyces cerevisiae, three of which are unpublished, provides new insight into the Mediator signaling network. In addition to the known modular organization of the Mediator subunits, MC EMiNEM reveals a hierarchical ordering of its internal information flow, which is putatively transmitted through structural changes within the complex. We identify the N-terminus of Med7 as a peripheral entity, entailing only local structural changes upon perturbation, while the C-terminus of Med7 and Med19 appear to play a central role. MC EMiNEM associates Mediator subunits to most directly affected genes, which, in conjunction with gene set enrichment analysis, allows us to construct an interaction map of Mediator subunits and transcription factors.