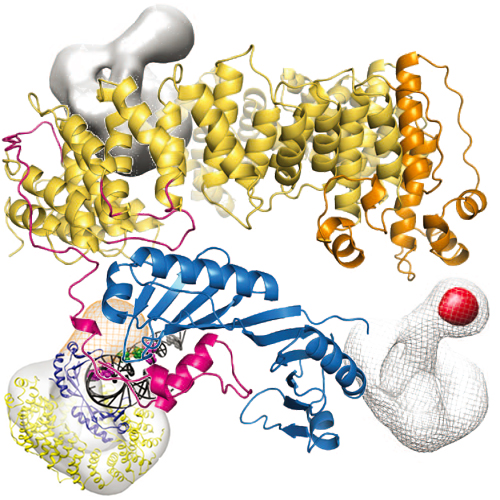
Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP
06-Jul-2011
Nature, 2011, doi:10.1038/nature10215, published on 06.07.2011
Swi2/Snf2-typeATPases regulate genome-associated processes such as transcription, replication and repair by catalysing the disruption, assembly or remodelling of nucleosomes or other protein–DNA complexes1,2. It has been suggested that ATP-driven motor activity along DNA disrupts target protein–DNA interactions in the remodelling reaction3–5. However, the complex and highly specific remodelling reactions are poorly understood, mostly because of a lack of high-resolution structural information about how remodellers bind to their substrate proteins. Mot1 (modifier of transcription 1 in Saccharomyces cerevisiae, denoted BTAF1 in humans) is a Swi2/Snf2 enzyme that specifically displaces the TATA box binding protein (TBP) from the promoter DNA and regulates transcription globally by generating a highly dynamic TBP pool in the cell6,7. As a Swi2/Snf2 enzyme that functions as a single polypeptide and interacts with a relatively simple substrate, Mot1 offers an ideal systemfromwhich to gain a better understanding of thisimportant enzyme family.Toreveal howMot1 specifically disrupts TBP–DNA complexes, we combined crystal and electron microscopy structures of Mot1–TBP from Encephalitozoon cuniculi with biochemical studies.Here weshowthatMot1 wraps around TBP and seems to act like a bottle opener: a spring-like array of 16 HEAT(huntingtin, elongation factor 3, protein phosphatase 2Aand lipid kinase TOR) repeats grips theDNA-distal side of TBP via loop insertions, and the Swi2/Snf2 domain binds to upstream DNA, positioned to weaken the TBP–DNAinteraction byDNAtranslocation. A‘latch’ subsequently blocks theDNA-binding groove ofTBP, acting as a chaperone to prevent DNA re-association and ensure efficient promoter clearance. This work shows how a remodelling enzyme can combine both motor and chaperone activities to achieve functional specificity using a conserved Swi2/Snf2 translocase.