Research Area E - Publications 2020
24-Nov-2020
Nat Commun 11, 5972 (2020), https://doi.org/10.1038/s41467-020-19603-1
Nat Commun., online article
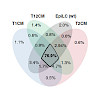
Genome-wide DNA demethylation is a unique feature of mammalian development and naïve pluripotent stem cells. Here, we describe a recently evolved pathway in which global hypomethylation is achieved by the coupling of active and passive demethylation. TET activity is required, albeit indirectly, for global demethylation, which mostly occurs at sites devoid of TET ...
21-Feb-2020
PLoS ONE, 15(2): e0229144, https://doi.org/10.1371/journal.pone.0229144
PLoS ONE, online article
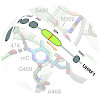
The multi-domain protein UHRF1 is essential for DNA methylation maintenance and binds DNA via a base-flipping mechanism with a preference for hemi-methylated CpG sites. We investigated its binding to hemi- and symmetrically modified DNA containing either 5-methylcytosine (mC), 5-hydroxymethylcytosine (hmC), 5-formylcytosine (fC), or 5-carboxylcytosine (caC). Our ...










