ANTICALIgN: visualizing, editing and analyzing combined nucleotide and amino acid sequence alignments for combinatorial protein engineering
30-May-2016
Protein Engineering, Design & Selection, vol. 29 no. 7, pp. 263–270, doi: 10.1093/protein/gzw016
Protein Engineering, Design & Selection, online article
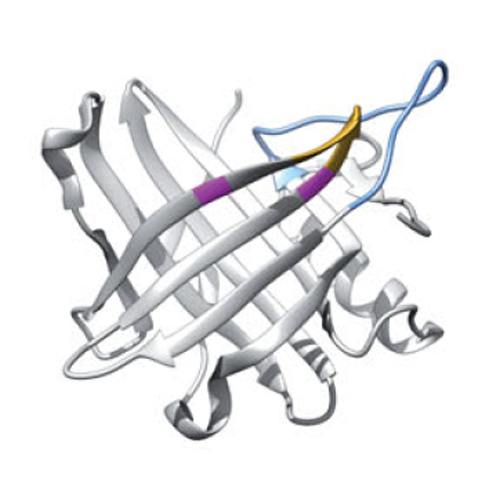
ANTICALIgN is an interactive software developed to simultaneously visualize, analyze and modify alignments of DNA and/or protein sequences that arise during combinatorial protein engineering, design and selection. ANTICALIgN combines powerful functions known from currently available sequence analysis tools with unique features for protein engineering, in particular the possibility to display and manipulate nucleotide sequences and their translated amino acid sequences at the same time. ANTICALIgN offers both template-based multiple sequence alignment (MSA), using the unmutated protein as reference, and conventional global alignment, to compare sequences that share an evolutionary relationship. The application of similarity-based clustering algorithms facilitates the identification of duplicates or of conserved sequence features among a set of selected clones. Imported nucleotide sequences from DNA sequence analysis are automatically translated into the corresponding amino acid sequences and displayed, offering numerous options for selecting reading frames, highlighting of sequence features and graphical layout of the MSA. The MSA complexity can be reduced by hiding the conserved nucleotide and/or amino acid residues, thus putting emphasis on the relevant mutated positions. ANTICALIgN is also able to handle suppressed stop codons or even to incorporate non-natural amino acids into a coding sequence. We demonstrate crucial functions of ANTICALIgN in an example of Anticalins selected from a lipocalin random library against the fibronectin extradomain B (ED-B), an established marker of tumor vasculature. Apart from engineered protein scaffolds, ANTICALIgN provides a powerful tool in the area of antibody engineering and for directed enzyme evolution.