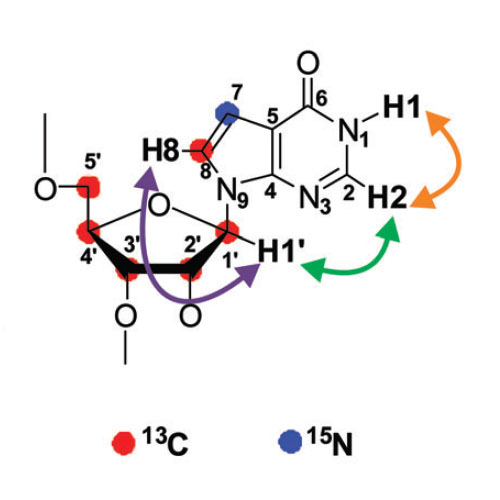
Site-Specific Isotope-Labeling of Inosine Phosphoramidites and NMR Analysis of an Inosine-Containing RNA Duplex
08-Sep-2016
Chem. Eur. J., 22, 1 – 11, DOI: 10.1002/chem.201602784
Chem. Eur. J., online article
Structural features and internal dynamics of inosine-containing RNAs are poorly understood. NMR studies of such RNAs require 13C,15N-labeling, which cannot be achieved using in vitro transcription as inosine and guanosine are not distinguished by RNA polymerase. Herein, we report the synthesis of an inosine phosphoramidite with selective 13C8 and 15N7-isotope incorporation in the base and uniform 13C-labeling of the ribose. Chemical synthesis of an RNA duplex containing four consecutive IU base pairs with this optimized isotope-labeling scheme greatly simplifies NMR spectra and resolves signal overlap. The absence of detectable NMR signals of imino protons and unusual inter-residue NOE correlations in this RNA indicate deviations from standard A-form geometry, consistent with reduced stability of this duplex seen in UV melting studies compared to its nonedited RNA counterparts. These studies indicate that the introduction of IU base pairs distorts and destabilizes RNA helices significantly compared to the also noncanonical GU base-pairs. Our optimized isotope-labeling scheme enables high-resolution NMR studies of inosine-edited RNAs.